Research
Neural circuit specialization
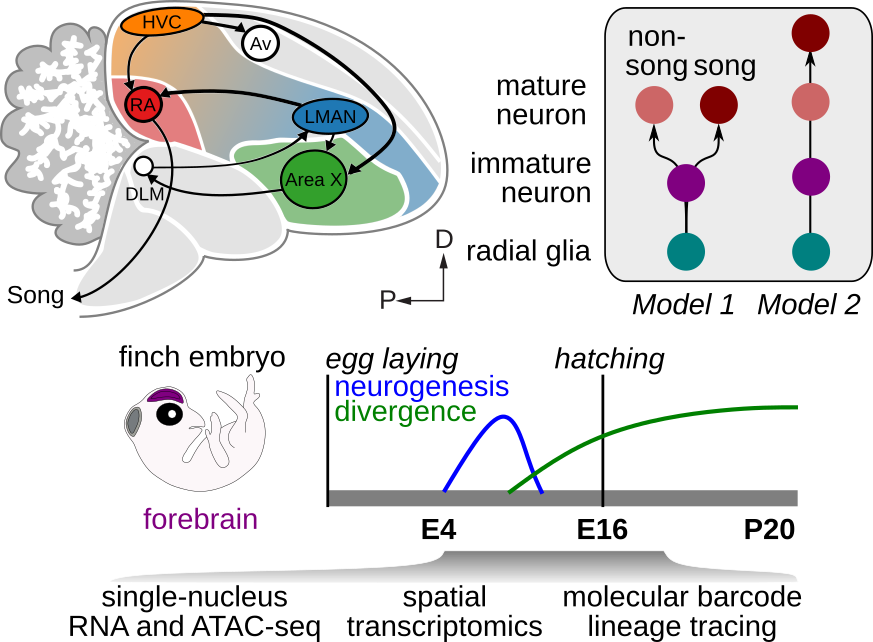
Complex learned behaviors require the coordinated activity of multiple specialized neuron types across interconnected brain regions. How do such circuits arise during development, and what makes them distinct from neighboring, unspecialized regions?
Birdsong is controlled by a dedicated set of brain nuclei — the song system — that is sharply demarcated from surrounding tissue and highly specialized for vocal-motor learning. This makes it an exceptional model for understanding how gene regulatory programs drive the emergence of circuit identity during development.
The lab uses cell-resolved molecular assays, spatial transcriptomics, and targeted gene manipulation in songbirds to decode the transcriptional logic that builds specialized circuits from a common developmental substrate.
Molecular mechanisms of motor skill stabilization
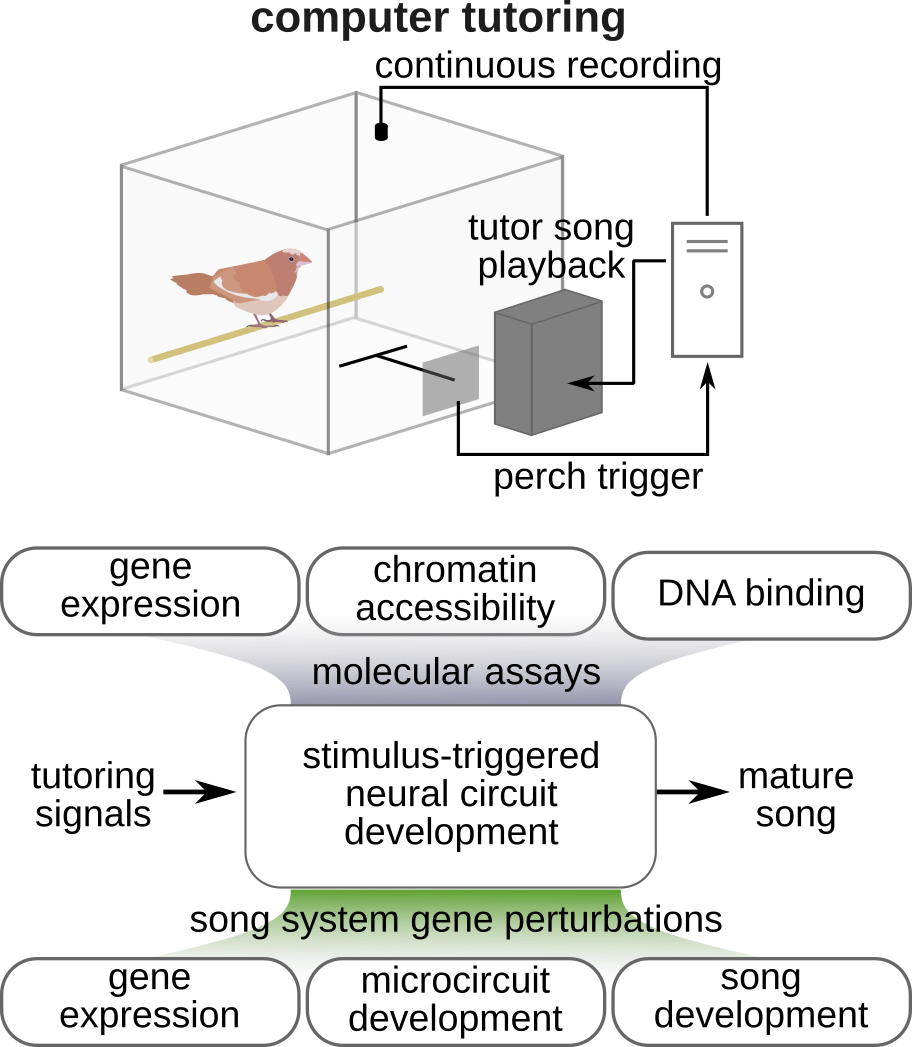
Like human speech, birdsong is learned during a sensitive period early in life. Young birds listen to an adult tutor, form a memory of the target song, and then practice until their own vocalizations converge on that template. This sensorimotor learning process is guided by a molecular program that opens a plastic window during development and then closes it as the circuit matures.
We study the molecular mechanisms that control this transition from plasticity to stability — asking which genes gate the sensitive period, how auditory experience modifies gene expression in vocal circuits, and whether these mechanisms can be leveraged to restore plasticity in adulthood.
Evolution of vocal motor circuits
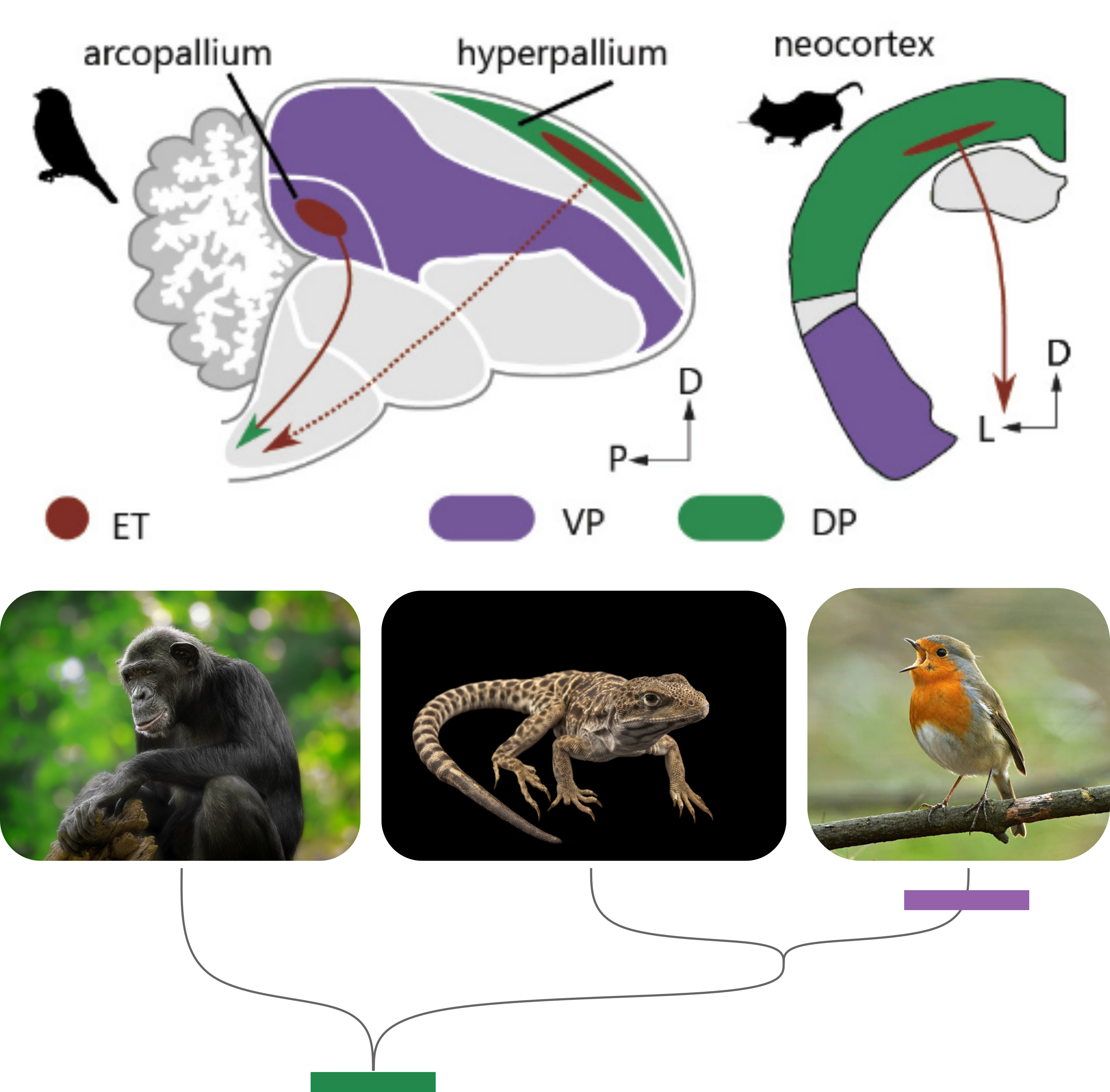
Vocal learning has evolved independently in songbirds, hummingbirds, parrots, cetaceans, and humans. Despite very different brain architectures, these lineages converge on remarkably similar circuit solutions — dedicated forebrain regions that connect auditory memory to motor output.
We compare the cellular composition, molecular identity, and developmental histories of vocal motor circuits across species and between birds and mammals. Our goal is to understand how similar neural circuits can arise from distinct developmental starting points, illuminating the deep logic of motor circuit evolution.

